Variant Annotations¶
Cases are processed via the bioinformatics pipeline and additional annotations are added by Cellbase. Once a case arrives to the DSS, additional annotations are added from resources that are currently unavailable in Cellbase.
| Annotation | Description | Source |
|---|---|---|
| gnomAD (exomes) | Population Germline Allele Frequency Database | Cellbase |
| Ensembl | Transcript and Protein ID annotations | Cellbase |
| RefSeq | Transcript annotations | Cellbase |
| ClinVar | Germline Database | Cellbase |
| NGTD | NHS resource | DSS |
| PanelApp | GEL resource | DSS |
| SV Frequencies | GEL dataset | DSS |
| CGC annotations | Somatic Database | DSS |
| Cancer Hotspots | Somatic Database | DSS |
| SVIG canonical variant list | Somatic Database | DSS |
| GEL 100K somatic frequencies | GEL dataset | DSS |
Info
Please note that COSMIC annotations have been removed from the DSS from release 6.18.0 onwards due to licensing changes. Please see Release notes for further details.
The Genome Aggregation Database (gnomAD) is a population frequency database of large-scale sequencing projects from healthy cohorts.
- Cellbase annotates whether a variant is present in the gnomAD exomes dataset.
- The DSS displays the total population germline allele frequency as well as the alternate allele frequency, alternate allele count, allele number, and number of individuals homozygous for the alternate allele in each subpopulation.
- The database version of gnomAD used by Cellbase is displayed in the DSS and may not always be the most recent. Please see the latest version of the Cancer Genome Analysis guide and Cellbase for further details.
The Cancer Hotspots Database consists of single residue and in-frame indel mutation hotspots identified in 24,592 tumor samples by the algorithm described in Chang et al. 2017 and Chang et al. 2016, as well as 164 hotspots reported in Bandlamudi et al., 2026.
Warning
Cancer hotspot annotations have not been updated to contain the latest 164 hotspots reported in Bandlamudi et al., 2026
- If the variant is located in a hotspot identified by Cancer Hotspots, the cancer hotspots icon will show in the database column in the variant grid.
- We currently match hotspots using the ensembl protein ID. Cancerhotspots.org have used Ensembl v75, while our pipeline uses MANE transcripts. Please see below for a comparison between all protein IDs.
- In cases where the protein ID does not match, we will not detect a hotspot in that gene. We recommend navigating to the cancerhotspots.org database via the linkout provided and manually searching for the hotspot, then comparing protein sequences to ensure the protein sequence matches at that position.
Proteins
| HGNC Symbol | Ensembl 75 | Ensembl 90 | MANE | Notes |
|---|---|---|---|---|
| ACVR1 | ENSP00000263640 | ENSP00000263640 | ENSP00000405004.1 | |
| AKT1 | ENSP00000270202 | ENSP00000450688 | ENSP00000497822.1 | |
| AKT3 | ENSP00000263826 | ENSP00000336943 | ENSP00000500582.1 | |
| ALK | ENSP00000373700 | ENSP00000373700 | ENSP00000373700.3 | |
| ANKRD11 | ENSP00000301030 | ENSP00000367581 | ENSP00000301030.4 | |
| APC | ENSP00000257430 | ENSP00000257430 | ENSP00000257430.4 | |
| AR | ENSP00000363822 | ENSP00000363822 | ENSP00000363822.3 | |
| ARAF | ENSP00000366244 | ENSP00000366244 | ENSP00000366244.4 | |
| ARID1A | ENSP00000320485 | ENSP00000320485 | ENSP00000320485.7 | |
| ARID1B | ENSP00000344546 | ENSP00000344546 | ENSP00000490491.2 | |
| ARID2 | ENSP00000335044 | ENSP00000335044 | ENSP00000335044.6 | |
| ASXL2 | ENSP00000391447 | ENSP00000391447 | ENSP00000391447.3 | |
| ATM | ENSP00000278616 | ENSP00000278616 | ENSP00000501606.1 | |
| ATRX | ENSP00000362441 | ENSP00000362441 | ENSP00000362441.4 | |
| AXIN1 | ENSP00000262320 | ENSP00000262320 | ENSP00000262320.3 | |
| AXL | ENSP00000301178 | ENSP00000301178 | ENSP00000301178.3 | |
| B2M | ENSP00000452780 | ENSP00000452780 | ENSP00000497910.1 | |
| BCL10 | ENSP00000359612 | ENSP00000359612 | ENSP00000498104.1 | |
| BCL2 | ENSP00000329623 | ENSP00000381185 | ENSP00000329623.3 | |
| BCL2L11 | ENSP00000376943 | ENSP00000482175 | ENSP00000376943.2 | |
| BCL6 | ENSP00000232014 | ENSP00000384371 | ENSP00000384371.2 | |
| BCOR | ENSP00000367705 | ENSP00000367705 | ENSP00000367705.4 | |
| BRAF | ENSP00000288602 | ENSP00000288602 | ENSP00000496776.1 | |
| BRCA2 | ENSP00000369497 | ENSP00000439902 | ENSP00000369497.3 | |
| BRD4 | ENSP00000263377 | ENSP00000263377 | ENSP00000506350.1 | |
| CARD11 | ENSP00000380150 | ENSP00000380150 | ENSP00000380150.4 | |
| CASP8 | ENSP00000351273 | ENSP00000264275 | ENSP00000501268.1 | |
| CBFB | ENSP00000415151 | ENSP00000415151 | ENSP00000415151.2 | |
| CBL | ENSP00000264033 | ENSP00000264033 | ENSP00000264033.3 | |
| CCND1 | ENSP00000227507 | ENSP00000227507 | ENSP00000227507.2 | |
| CCND3 | ENSP00000362082 | ENSP00000362082 | ENSP00000362082.5 | |
| CD79B | ENSP00000376544 | ENSP00000376544 | ENSP00000006750.4 | |
| CDH1 | ENSP00000261769 | ENSP00000261769 | ENSP00000261769.4 | |
| CDK12 | ENSP00000398880 | ENSP00000398880 | ENSP00000398880.4 | |
| CDK4 | ENSP00000257904 | ENSP00000257904 | ENSP00000257904.5 | |
| CDK6 | ENSP00000265734 | ENSP00000265734 | ENSP00000397087.3 | |
| CDKN1A | ENSP00000244741 | ENSP00000409259 | ENSP00000244741.6 | |
| CDKN1B | ENSP00000228872 | ENSP00000228872 | ENSP00000228872.4 | |
| CDKN2A | ENSP00000307101 | ENSP00000307101 | ENSP00000307101.5 | |
| CHEK2 | ENSP00000329178 | ENSP00000329178 | ENSP00000385747.1 | |
| CIC | ENSP00000458663 | ENSP00000458663 | ENSP00000505728.1 | |
| CREBBP | ENSP00000262367 | ENSP00000262367 | ENSP00000262367.5 | |
| CRLF2 | ENSP00000370978 | ENSP00000383641 | ENSP00000383641.3 | |
| CTCF | ENSP00000264010 | ENSP00000264010 | ENSP00000264010.4 | |
| CTLA4 | ENSP00000303939 | ENSP00000303939 | ENSP00000497102.1 | |
| CTNNB1 | ENSP00000344456 | ENSP00000344456 | ENSP00000344456.5 | |
| CUL3 | ENSP00000264414 | ENSP00000264414 | ENSP00000264414.4 | |
| CYSLTR2 | ENSP00000282018 | ENSP00000282018 | ENSP00000508181.1 | |
| DICER1 | ENSP00000343745 | ENSP00000343745 | ENSP00000343745.3 | |
| DIS3 | ENSP00000366997 | ENSP00000366997 | ENSP00000366997.4 | |
| DNAJB1 | ENSP00000254322 | ENSP00000254322 | ENSP00000254322.1 | |
| DNMT1 | ENSP00000345739 | ENSP00000352516 | ENSP00000352516.3 | |
| DNMT3A | ENSP00000264709 | ENSP00000324375 | ENSP00000324375.5 | |
| DNMT3B | ENSP00000328547 | ENSP00000328547 | ENSP00000328547.2 | |
| EGFR | ENSP00000275493 | ENSP00000275493 | ENSP00000275493.2 | |
| EIF1AX | ENSP00000368927 | ENSP00000368927 | ENSP00000368927.5 | |
| EIF4A2 | ENSP00000326381 | ENSP00000326381 | ENSP00000326381.5 | |
| EP300 | ENSP00000263253 | ENSP00000263253 | ENSP00000263253.7 | |
| EPAS1 | ENSP00000263734 | ENSP00000263734 | ENSP00000263734.3 | |
| EPHA3 | ENSP00000337451 | ENSP00000337451 | ENSP00000337451.2 | |
| EPHA7 | ENSP00000358309 | ENSP00000358309 | ENSP00000358309.4 | |
| EPHB1 | ENSP00000381097 | ENSP00000381097 | ENSP00000381097.3 | |
| ERBB2 | ENSP00000269571 | ENSP00000269571 | ENSP00000269571.4 | |
| ERBB3 | ENSP00000267101 | ENSP00000267101 | ENSP00000267101.4 | |
| ERBB4 | ENSP00000342235 | ENSP00000342235 | ENSP00000342235.4 | |
| ERCC2 | ENSP00000375809 | ENSP00000375809 | ENSP00000375809.4 | |
| ERCC3 | ENSP00000285398 | ENSP00000285398 | ENSP00000285398.2 | |
| ERG | ENSP00000288319 | ENSP00000414150 | ENSP00000288319.7 | |
| ERRFI1 | ENSP00000366702 | ENSP00000366702 | ENSP00000366702.5 | |
| ESR1 | ENSP00000206249 | ENSP00000206249 | ENSP00000206249.3 | |
| ETV6 | ENSP00000379658 | ENSP00000379658 | ENSP00000379658.3 | |
| EZH2 | ENSP00000320147 | ENSP00000320147 | ENSP00000320147.2 | |
| FAM58A | ENSP00000384396 | ENSP00000461135 | ENSP00000461135.1 | CCNQ in MANE |
| FAT1 | ENSP00000406229 | ENSP00000406229 | ENSP00000406229.2 | |
| FBXW7 | ENSP00000281708 | ENSP00000281708 | ENSP00000281708.3 | |
| FGFR1 | ENSP00000393312 | ENSP00000400162 | ENSP00000400162.2 | |
| FGFR2 | ENSP00000351276 | ENSP00000410294 | ENSP00000351276.6 | |
| FGFR3 | ENSP00000260795 | ENSP00000414914 | ENSP00000414914.2 | |
| FGFR4 | ENSP00000292408 | ENSP00000292408 | ENSP00000292408.4 | |
| FH | ENSP00000355518 | ENSP00000355518 | ENSP00000355518.4 | |
| FLT3 | ENSP00000241453 | ENSP00000241453 | ENSP00000241453.7 | |
| FOXA1 | ENSP00000250448 | ENSP00000250448 | ENSP00000250448.3 | |
| FOXL2 | ENSP00000333188 | ENSP00000333188 | ENSP00000497217.1 | |
| FOXO1 | ENSP00000368880 | ENSP00000368880 | ENSP00000368880.4 | |
| FOXP1 | ENSP00000318902 | ENSP00000318902 | ENSP00000497369.1 | |
| FUBP1 | ENSP00000359804 | ENSP00000359804 | ENSP00000359804.2 | |
| GATA2 | ENSP00000345681 | ENSP00000345681 | ENSP00000345681.2 | |
| GATA3 | ENSP00000341619 | ENSP00000368632 | ENSP00000368632.3 | |
| GLI1 | ENSP00000228682 | ENSP00000228682 | ENSP00000228682.2 | |
| GNA11 | ENSP00000078429 | ENSP00000078429 | ENSP00000078429.3 | |
| GNAQ | ENSP00000286548 | ENSP00000286548 | ENSP00000286548.4 | |
| GNAS | ENSP00000360126 | ENSP00000346328 | ENSP00000360115.3 | |
| GPS2 | ENSP00000370104 | ENSP00000379841 | ENSP00000370104.2 | |
| GSK3B | ENSP00000324806 | ENSP00000324806 | ENSP00000264235.9 | |
| GTF2I | ENSP00000322542 | ENSP00000460070 | ENSP00000460070.1 | |
| H3F3A | ENSP00000355778 | ENSP00000355780 | ENSP00000355780 | |
| HGF | ENSP00000222390 | ENSP00000222390 | ENSP00000222390.5 | |
| HIST1H1C | ENSP00000339566 | ENSP00000339566 | ENSP00000339566.3 | H1-2 in MANE |
| HIST1H3B | ENSP00000244661 | ENSP00000484841 | ENSP00000484841.2 | H3-C2 in MANE |
| HIST1H3C | ENSP00000439493 | ENSP00000484658 | ENSP00000484658.2 | H3-C3 in MANE |
| HIST1H3H | ENSP00000358160 | ENSP00000358160 | ENSP00000509120.1 | H3C10 in MANE |
| HLA-A | ENSP00000366005 | ENSP00000366002 | ENSP00000366005.5 | |
| HLA-B | ENSP00000399168 | ENSP00000399168 | ENSP00000399168.2 | |
| HNF1A | ENSP00000257555 | ENSP00000257555 | ENSP00000257555.5 | |
| HRAS | ENSP00000309845 | ENSP00000407586 | ENSP00000309845.7 | |
| IDH1 | ENSP00000260985 | ENSP00000260985 | ENSP00000260985.2 | |
| IDH2 | ENSP00000331897 | ENSP00000331897 | ENSP00000331897.4 | |
| IGF1R | ENSP00000268035 | ENSP00000268035 | ENSP00000497069.1 | |
| IKZF1 | ENSP00000331614 | ENSP00000331614 | ENSP00000331614.3 | |
| IL7R | ENSP00000306157 | ENSP00000306157 | ENSP00000306157.3 | |
| INPPL1 | ENSP00000298229 | ENSP00000298229 | ENSP00000298229.2 | |
| IRF4 | ENSP00000370343 | ENSP00000370343 | ENSP00000370343.4 | |
| IRS2 | ENSP00000365016 | ENSP00000365016 | ENSP00000365016.3 | |
| JAK1 | ENSP00000343204 | ENSP00000343204 | ENSP00000343204.4 | |
| JUN | ENSP00000360266 | ENSP00000360266 | ENSP00000360266.2 | |
| KDM6A | ENSP00000367203 | ENSP00000367203 | ENSP00000483595.2 | |
| KDR | ENSP00000263923 | ENSP00000263923 | ENSP00000263923.4 | |
| KEAP1 | ENSP00000171111 | ENSP00000377245 | ENSP00000171111.4 | |
| KIT | ENSP00000288135 | ENSP00000288135 | ENSP00000288135.6 | |
| KLF4 | ENSP00000363804 | ENSP00000363804 | ENSP00000363804.4 | |
| KMT2A | ENSP00000436786 | ENSP00000436786 | ENSP00000436786.2 | |
| KMT2C | ENSP00000262189 | ENSP00000262189 | ENSP00000262189.6 | |
| KMT2D | ENSP00000301067 | ENSP00000301067 | ENSP00000301067.7 | |
| KNSTRN | ENSP00000249776 | ENSP00000249776 | ENSP00000249776.8 | |
| KRAS | ENSP00000256078 | ENSP00000256078 | ENSP00000308495.3 | |
| LYN | ENSP00000428924 | ENSP00000428924 | ENSP00000428924.1 | |
| MAP2K1 | ENSP00000302486 | ENSP00000302486 | ENSP00000302486.5 | |
| MAP2K2 | ENSP00000262948 | ENSP00000262948 | ENSP00000262948.4 | |
| MAP2K4 | ENSP00000262445 | ENSP00000410402 | ENSP00000262445.5 | |
| MAP3K1 | ENSP00000382423 | ENSP00000382423 | ENSP00000382423.3 | |
| MAP3K13 | ENSP00000265026 | ENSP00000265026 | ENSP00000265026.3 | |
| MAPK1 | ENSP00000215832 | ENSP00000215832 | ENSP00000215832.7 | |
| MAX | ENSP00000351490 | ENSP00000351490 | ENSP00000351490.4 | |
| MED12 | ENSP00000363193 | ENSP00000363193 | ENSP00000363193.3 | |
| MEF2B | ENSP00000162023 | ENSP00000402154 | ENSP00000402154.2 | |
| MET | ENSP00000380860 | ENSP00000317272 | ENSP00000380860.3 | |
| MGA | ENSP00000219905 | ENSP00000219905 | N/A | non-coding in MANE |
| MSH2 | ENSP00000233146 | ENSP00000233146 | ENSP00000233146.2 | |
| MST1 | ENSP00000414287 | ENSP00000414287 | ENSP00000414287.2 | |
| MTOR | ENSP00000354558 | ENSP00000354558 | ENSP00000354558.4 | |
| MYC | ENSP00000367207 | ENSP00000479618 | ENSP00000478887.2 | |
| MYCN | ENSP00000281043 | ENSP00000281043 | ENSP00000281043.3 | |
| MYD88 | ENSP00000379625 | ENSP00000379625 | ENSP00000498360.2 | |
| MYOD1 | ENSP00000250003 | ENSP00000250003 | ENSP00000250003.3 | |
| NCOA3 | ENSP00000361066 | ENSP00000361066 | ENSP00000361066.3 | |
| NF1 | ENSP00000351015 | ENSP00000351015 | ENSP00000351015.4 | |
| NFE2L2 | ENSP00000380252 | ENSP00000380252 | ENSP00000380252.3 | |
| NOTCH1 | ENSP00000277541 | ENSP00000277541 | ENSP00000498587.1 | |
| NOTCH2 | ENSP00000256646 | ENSP00000256646 | ENSP00000256646.2 | |
| NRAS | ENSP00000358548 | ENSP00000358548 | ENSP00000358548.4 | |
| NSD1 | ENSP00000395929 | ENSP00000395929 | ENSP00000395929.2 | |
| NUP93 | ENSP00000310668 | ENSP00000310668 | ENSP00000310668.5 | |
| PAK7 | ENSP00000322957 | ENSP00000367686 | ENSP00000322957.5 | PAK5 in MANE |
| PARP1 | ENSP00000355759 | ENSP00000355759 | ENSP00000355759.5 | |
| PAX5 | ENSP00000350844 | ENSP00000350844 | ENSP00000350844.4 | |
| PBRM1 | ENSP00000378307 | ENSP00000378307 | N/A | Pending MANE review |
| PDGFRA | ENSP00000257290 | ENSP00000257290 | ENSP00000257290.5 | |
| PDPK1 | ENSP00000344220 | ENSP00000344220 | ENSP00000344220.4 | |
| PGR | ENSP00000325120 | ENSP00000325120 | ENSP00000325120.5 | |
| PIK3CA | ENSP00000263967 | ENSP00000263967 | ENSP00000263967.3 | |
| PIK3CB | ENSP00000289153 | ENSP00000289153 | ENSP00000501150.1 | |
| PIK3CD | ENSP00000366563 | ENSP00000366563 | ENSP00000366563.4 | |
| PIK3R1 | ENSP00000274335 | ENSP00000428056 | ENSP00000428056.1 | |
| PIK3R2 | ENSP00000222254 | ENSP00000222254 | ENSP00000222254.6 | |
| PIM1 | ENSP00000362608 | ENSP00000362608 | ENSP00000362608.5 | |
| PLK2 | ENSP00000274289 | ENSP00000274289 | ENSP00000274289.3 | |
| PMS2 | ENSP00000265849 | ENSP00000265849 | ENSP00000265849.7 | |
| POLE | ENSP00000322570 | ENSP00000322570 | ENSP00000322570.5 | |
| PPM1D | ENSP00000306682 | ENSP00000306682 | ENSP00000306682.2 | |
| PPP2R1A | ENSP00000324804 | ENSP00000324804 | ENSP00000324804.6 | |
| PPP4R2 | ENSP00000432104 | ENSP00000349124 | ENSP00000349124.5 | |
| PPP6C | ENSP00000362648 | ENSP00000392147 | ENSP00000362648.4 | |
| PRDM14 | ENSP00000276594 | ENSP00000276594 | ENSP00000276594.2 | |
| PREX2 | ENSP00000288368 | ENSP00000288368 | ENSP00000288368.4 | |
| PRKCI | ENSP00000295797 | ENSP00000295797 | ENSP00000295797.4 | |
| PTEN | ENSP00000361021 | ENSP00000361021 | ENSP00000361021.3 | |
| PTPN11 | ENSP00000340944 | ENSP00000340944 | ENSP00000340944.3 | |
| PTPRD | ENSP00000348812 | ENSP00000370593 | ENSP00000370593.3 | |
| PTPRS | ENSP00000349932 | ENSP00000465185 | ENSP00000262963.8 | |
| PTPRT | ENSP00000362294 | ENSP00000362294 | ENSP00000362283.1 | |
| RAC1 | ENSP00000348461 | ENSP00000348461 | ENSP00000258737.7 | |
| RAD50 | ENSP00000265335 | ENSP00000368100 | ENSP00000368100.4 | |
| RAD51C | ENSP00000336701 | ENSP00000336701 | ENSP00000336701.4 | |
| RAD52 | ENSP00000351284 | ENSP00000351284 | ENSP00000351284.3 | |
| RAF1 | ENSP00000251849 | ENSP00000251849 | ENSP00000251849.4 | |
| RARA | ENSP00000254066 | ENSP00000254066 | ENSP00000254066.5 | |
| RB1 | ENSP00000267163 | ENSP00000267163 | ENSP00000267163.4 | |
| RBM10 | ENSP00000328848 | ENSP00000366829 | ENSP00000366829.3 | |
| RECQL4 | ENSP00000475456 | ENSP00000482313 | ENSP00000482313.2 | |
| RET | ENSP00000347942 | ENSP00000347942 | ENSP00000347942.3 | |
| RHEB | ENSP00000262187 | ENSP00000262187 | ENSP00000262187.5 | |
| RHOA | ENSP00000400175 | ENSP00000400175 | ENSP00000400175.1 | |
| RICTOR | ENSP00000349959 | ENSP00000349959 | ENSP00000349959.3 | |
| RIT1 | ENSP00000357306 | ENSP00000357306 | ENSP00000357306.3 | |
| RNF43 | ENSP00000385328 | ENSP00000462764 | ENSP00000385328.2 | |
| RPS6KA4 | ENSP00000333896 | ENSP00000333896 | ENSP00000333896.4 | |
| RPTOR | ENSP00000307272 | ENSP00000307272 | ENSP00000307272.3 | |
| RRAS2 | ENSP00000256196 | ENSP00000437547 | ENSP00000256196.4 | |
| RUNX1 | ENSP00000300305 | ENSP00000409227 | ENSP00000501943.1 | |
| RXRA | ENSP00000419692 | ENSP00000419692 | ENSP00000419692.1 | |
| SDHA | ENSP00000264932 | ENSP00000264932 | ENSP00000264932.6 | |
| SDHAF2 | ENSP00000301761 | ENSP00000301761 | ENSP00000301761.3 | |
| SESN2 | ENSP00000253063 | ENSP00000253063 | ENSP00000253063.3 | |
| SETD2 | ENSP00000386759 | ENSP00000386759 | ENSP00000386759.3 | |
| SF3B1 | ENSP00000335321 | ENSP00000335321 | ENSP00000335321.6 | |
| SMAD2 | ENSP00000262160 | ENSP00000349282 | ENSP00000262160.6 | |
| SMAD3 | ENSP00000332973 | ENSP00000332973 | ENSP00000332973.4 | |
| SMAD4 | ENSP00000341551 | ENSP00000341551 | ENSP00000341551.3 | |
| SMARCA4 | ENSP00000343896 | ENSP00000397783 | ENSP00000343896.4 | |
| SMARCB1 | ENSP00000263121 | ENSP00000263121 | ENSP00000494049.2 | |
| SMARCD1 | ENSP00000378414 | ENSP00000378414 | ENSP00000378414.4 | |
| SMO | ENSP00000249373 | ENSP00000249373 | ENSP00000249373.3 | |
| SOCS1 | ENSP00000329418 | ENSP00000329418 | ENSP00000329418.2 | |
| SOS1 | ENSP00000384675 | ENSP00000384675 | ENSP00000384675.2 | |
| SOX17 | ENSP00000297316 | ENSP00000297316 | ENSP00000297316.4 | |
| SPOP | ENSP00000240327 | ENSP00000377001 | ENSP00000425905.1 | |
| SPRED1 | ENSP00000299084 | ENSP00000299084 | ENSP00000299084.4 | |
| SRSF2 | ENSP00000353089 | ENSP00000376276 | ENSP00000353089.5 | |
| STAG2 | ENSP00000218089 | ENSP00000218089 | ENSP00000360187.4 | |
| STAT3 | ENSP00000264657 | ENSP00000264657 | ENSP00000264657.4 | |
| STK11 | ENSP00000324856 | ENSP00000324856 | ENSP00000324856.6 | |
| STK19 | ENSP00000364480 | ENSP00000364482 | ENSP00000509445.1 | |
| SUZ12 | ENSP00000316578 | ENSP00000316578 | ENSP00000316578.5 | |
| TBX3 | ENSP00000257566 | ENSP00000257566 | ENSP00000257567.2 | |
| TCEB1 | ENSP00000284811 | ENSP00000428171 | ENSP00000428171.1 | ELOC in MANE |
| TCF3 | ENSP00000344375 | ENSP00000262965 | ENSP00000262965.5 | |
| TCF7L2 | ENSP00000444972 | ENSP00000358404 | ENSP00000348274.4 | |
| TET1 | ENSP00000362748 | ENSP00000362748 | ENSP00000362748.4 | |
| TET2 | ENSP00000369351 | ENSP00000369351 | ENSP00000369351.4 | |
| TGFBR1 | ENSP00000364133 | ENSP00000364133 | ENSP00000364133.4 | |
| TGFBR2 | ENSP00000351905 | ENSP00000351905 | ENSP00000295754.5 | |
| TNFRSF14 | ENSP00000347948 | ENSP00000347948 | ENSP00000347948.4 | |
| TP53 | ENSP00000269305 | ENSP00000269305 | ENSP00000269305.4 | |
| TP63 | ENSP00000264731 | ENSP00000264731 | ENSP00000264731.3 | |
| TRAF7 | ENSP00000318944 | ENSP00000318944 | ENSP00000318944.6 | |
| TSC1 | ENSP00000298552 | ENSP00000298552 | ENSP00000298552.3 | |
| U2AF1 | ENSP00000291552 | ENSP00000291552 | ENSP00000291552.4 | |
| VHL | ENSP00000256474 | ENSP00000256474 | ENSP00000256474.3 | |
| WHSC1 | ENSP00000372347 | ENSP00000423972 | ENSP00000423972.1 | NSD2 in MANE |
| XPO1 | ENSP00000384863 | ENSP00000384863 | ENSP00000384863.2 |
Info
Cancer hotspot annotations are not currently available for splice variants or InDels, but will be in a future release. Until then, we recommend you check the cancerhotspots.org website for these variants.
We annotate variants against the SVIG canonical variant list. We display the SVIG flag in the cDSS for two scenarios:
- Exact match: if the exact variant is found in the SVIG canonical variant list, we display an 'SVIG' flag with the associated tooltip "This is a variant from the Somatic Variant Interpretation Guidelines (SVIG) canonical variant list"
- Region overlap: if the variant falls into the genomic region where an SVIG canonical variant is located, we display an 'SVIG region' flag and the associated tooltip "This is a variant falling into the genomic region where an SVIG canonical variant is located"
We annotate variants against the GEL 100K somatic frequencies dataset derived from the data published by Sonsinsky et al., 2024 encompassing a total of 11,374 samples. The 100K somatic frequencies is a large, accurate and robust dataset from the UK population which is ideal for case counting. At present, the dataset contains only small non-synonymous somatic variants from solid tumours.
Variant Grid¶
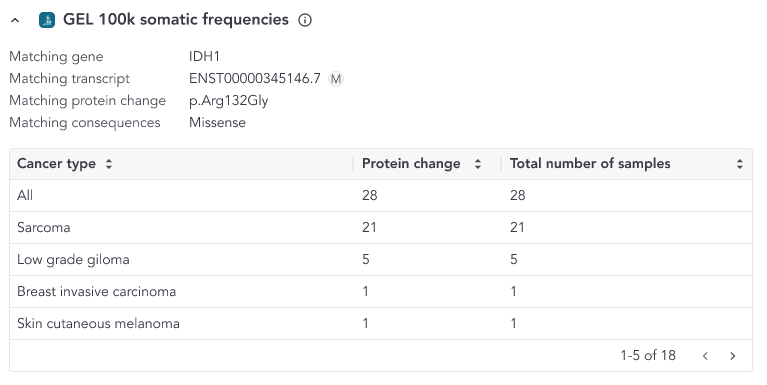
We display the GEL 100K somatic frequencies annotation in the variant grid across three scenarios:
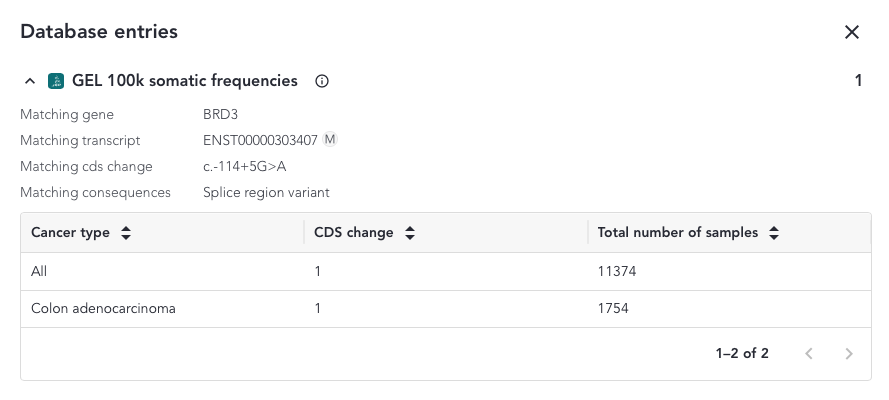
- Gene match: if the variant is annotated to a gene that was included in the GEL 100K somatic frequencies
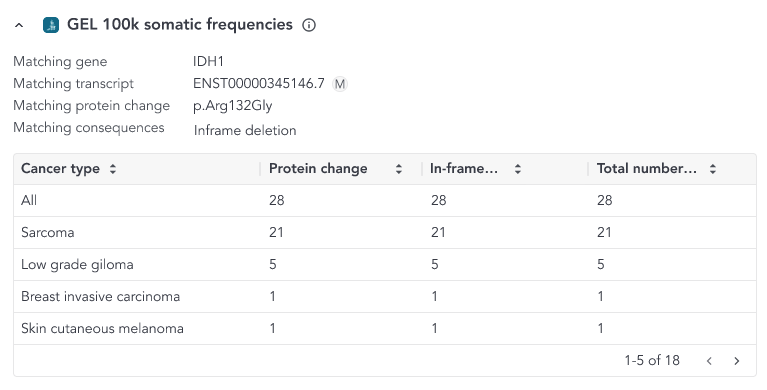
- Protein match: if the variant results in a matching protein sequence change, without matching the coding sequence (nucleotide) change
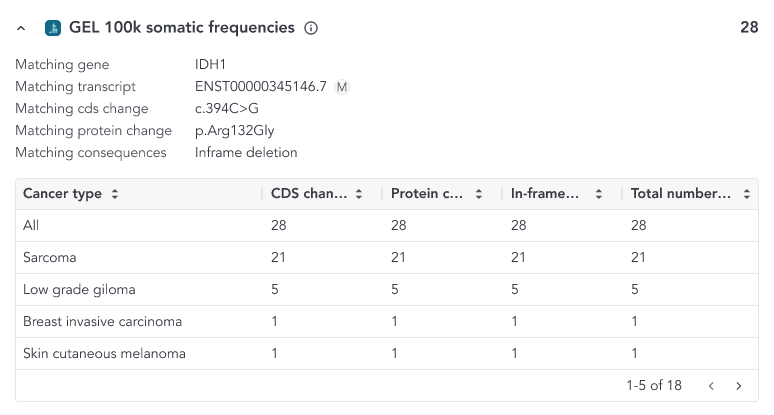
- Coding sequence match: if the variant matches at the coding sequence (nucleotide) and protein sequence change level
Each match is represented differently in the variant grid as shown below.
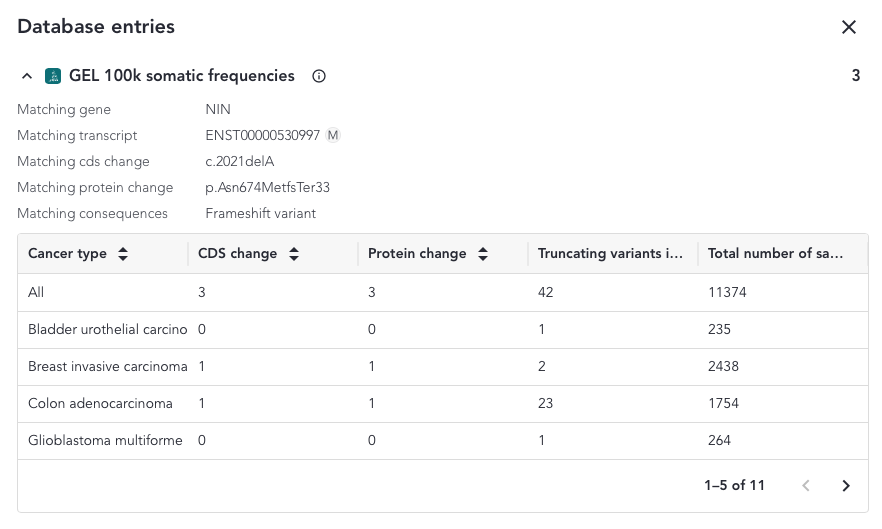
Database Entries Overlay¶
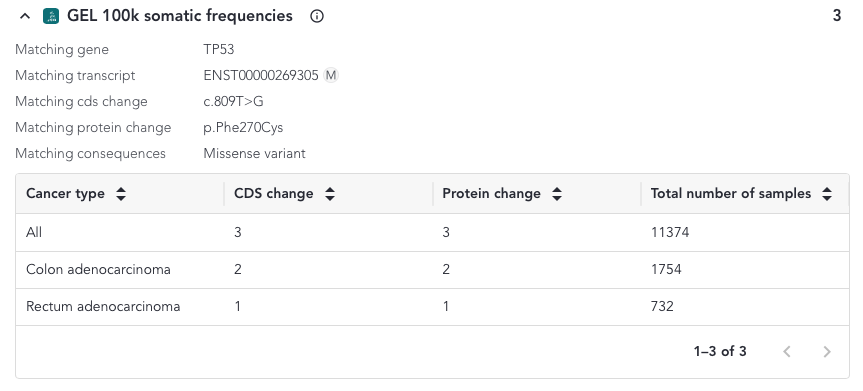
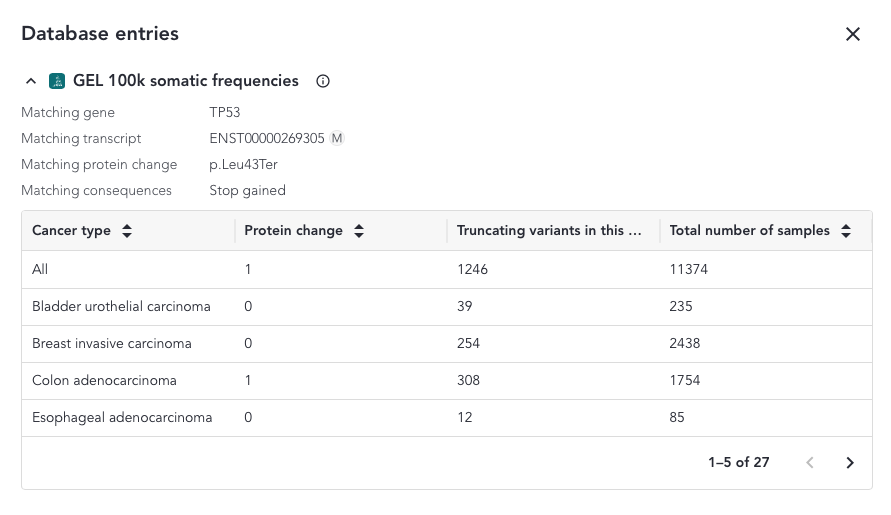
When a protein or a coding sequence match is found, the overlay includes a table where a breakdown of all the samples and cancer sample types included in the study that match the variant is shown.
The content of the overlay depends on the match and the variant type and can be accessed from the somatic small variant grid. The possible column defintions are shown below;
- Cancer Type - All the tumour types listed in alphabetical order
- CDS change - Number of samples with the searched variant resulting in the same coding sequence change
- Protein change - Number of samples with the searched variant resulting in the same protein change
- Truncating variants in this position and downstream - Counts for all truncating variants at the same position of further downstream ll
- In-frame deletions (within the same amino acid region) - Counts for all in-frame deletions within the same amino acid region
- Total number of samples - Total number of samples with that tumour type in the data set
Sample counts = The number of samples per cancer type and across all samples included in the study
Ensembl is a genome browser that provides gene annotations.
- Cellbase annotates each variant with a transcript and protein ID from Ensembl.
- The DSS displays these annotations in the variant list and variant details page.
- The database version of Ensembl used by Cellbase may not always be the most recent. Please see the latest version of the Cancer Genome Analysis guide and Cellbase for further details.
RefSeq is a sequence annotation resource.
- Transcripts included in the MANE project will have a RefSeq transcript ID annotation that corresponds to the Ensembl transcript ID.
- The DSS displays these annotations in the variant list overlay and variant details page.
ClinVar is a public database of variant and phenotype relationships alongside supporting evidence and clinical interpretation histories.
- Cellbase annotates whether a variant is present in Clinvar.
- The DSS displays the ClinVar ID, interpretation, review status and number of entries alongside a link to the ClinVar record in the variant list and variant details page for germline variants.
- The database version of ClinVar used by Cellbase is displayed in the DSS and may not always be the most recent. Please see the latest version of the Cancer Genome Analysis guide and Cellbase for further details.
The NGTD for cancer specifies which genomic tests are commissioned by the NHS in England, the technology by which they are available, and the patients who will be eligible to access to a test.
The DSS displays NGTD annotations in the tooltips of the genes shown in the variant grid and the gene information section of the variant details page.
NGTD annotations in the DSS depend on clinical indication groups. If the affected gene for a variant is present in the NGTD and is a target for any clinical indication within the same group as the patient's clinical indication (regardless of variant type), then the variant is annotated with the clinical indication group (e.g., Solid tumours) followed by all the clinical indication codes where the affected gene is a target (e.g., M1). If the affected gene is present in the NGTD but is a target for a different clinical indication group, the variant is simply annotated as present in the NGTD.
Info
The publication date of the NGTD file used is missing in the variant lists and detail pages of all cases ingested into the DSS between the 6.11.0 release (23/07/2025) and the 6.14.0 release (03/12/2025), please refer to the Cancer Genome Analysis guide for further details. The issue has been fixed as of the 6.14.0 release (03/12/2025) and any cases arriving to the DSS from this point onwards will be unaffected.
The GeL PanelApp knowledgebase allows virtual gene panels related to human disorders to be created, stored and queried and is used as the platform for achieving consensus on gene panels in the NHS Genomic Medicine Service (GMS).
- The Ensembl gene ID of each variant is used to match the variant to Panels in PanelApp.
- The Panels in which the affected gene is present are then shown on the variant list and variant details page.
Population germline allele frequency for the breakpoints of a given structural variant based on a panel of normals, which consists of germline structural variants observed in an internal Genomics England dataset of about 6,000 samples from unrelated individuals. If a variant has two breakpoints, maximal value of allele frequency among the two is reported. Not found suggests that this variant was not seen in the Genomics England dataset. If an SV is found to overlap with any SVs identified in the internal panels of normals, the population frequency will be annotated to the variant and displayed on the variant list and variant details page.
The Cancer Gene Census comprises a catalogue of genes which contain variants that have been causally implicated in cancer. In the Cancer DSS, we use the CGC database to annotate whether a gene is an oncogene, a tumour suppressor gene or both i.e., the role of the gene is ambiguous. For further information see Nature Reviews.
- The affected gene of each variant is compared against the CGC data.
- If a match is found, the annotation is attached to the affected gene and displayed in the DSS.